Testing antibiotic resistance with a fast, cheap, and easy method
“We have developed a technique in our laboratories that allows us to obtain an antibiogram within 2-4 hours – instead of the current 24 hours for the most common germs and one month for tuberculosis,” says Dr Sandor Kasas at EPFL. Professor Ronnie Willaert at Vrije Universiteit Brussel adds: “Our technique is not only faster but also simpler and much cheaper than all those existing now.”
Antibiotic resistance happens when bacteria develop the ability to defeat the drugs designed to kill them. It has now grown into a global public health issue. It was responsible for at least 1.27 million deaths worldwide in 2019 while being involved in nearly five million deaths. Every year, the US sees almost three million antimicrobial-resistant infections, with the cost of treating the six most common ones at over $4.6 billion. The EU sees almost 700,000 cases each year, which cost it an estimated €1.5 billion.
Antibiotic sensitivity testing (AST) uses culture methods that expose bacteria to antibiotics, or genetic methods to determine if bacteria possesses genes that confer resistance. Typical ASTs last up to 24 hours or even longer for slow-growing bacteria – a timeframe that can mean life or death in a clinical setting. There have been some faster ASTs developed in recent years, but they tend to be complex, needing sophisticated and expensive equipment.
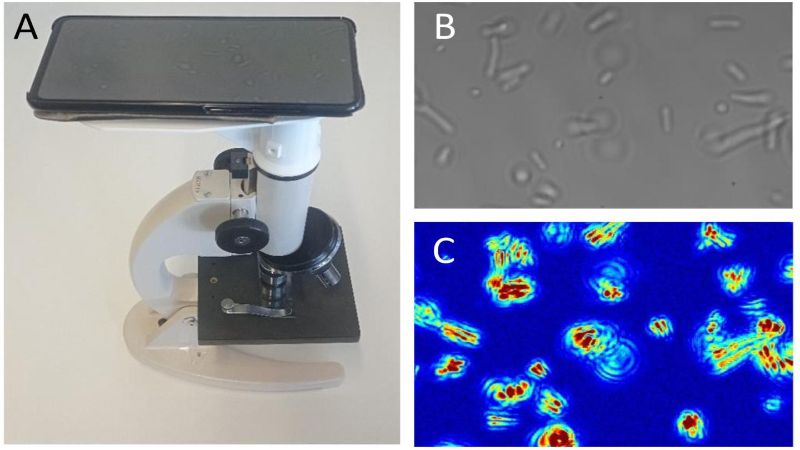
Now, researchers led by Kasas and Willaert have developed a fast, cheap, and widely accessible method based on optical microscopy that can perform an AST with single-cell sensitivity and without needing to attach or label bacteria. The technique uses a basic, conventional optical microscope, a camera or mobile phone, and dedicated software. The joint research project is published in PNAS.
The new technique is called optical nanomotion detection (ONMD), and involves the monitoring of nanoscale vibrations of single bacterial before and while being exposed to antibiotics. The monitoring is performed with a basic optical microscope, a video camera or a mobile phone.
The ONMD technique monitors the microscopic oscillations of bacterial cells (nanomotion) that characterize living organism and can be considered as a “signature of life”. Indeed, nanomotion lasts as long the organism is alive but stops immediately when it is dead. In the ONMD technique, bacterial nanomotion is recorded in a movie in which all individual cell displacements are monitored with sub-pixel resolution.
The researchers used ONMD to successfully detect the sensitivity of numerous bacteria to antibiotics. Escherichia coli, Staphylococcus aureus, Lactobacillus rhamnosus, and Mycobacterium smegmatis (a non-pathogenic bacterial model for tuberculosis) sensitivities to the antibiotics ampicillin, streptomycin, doxycycline, and vancomycin was determined in less than two hours.
The ONMD not only monitors the bacteria life-death transitions upon exposure to different antibiotics but also highlights changes in the bacteria’s metabolism caused by the availability of nutrients. The tests showed that ONMD can assess the sensitivity or resistance of bacterial cells to antibiotics in a simple and rapid way by monitoring cellular oscillations.
The authors state: “The simplicity and efficiency of the method make it a game-changer in the field of AST” as it can be applied to a wide range of bacteria, which has significant implications for clinical and research applications.